Biological Modelling Simulation ...
Year: 2005 - 2006
Programming Language & Tools: Microsoft .NET C#
Title: Biological Cell Model On Grid Computing
Simulation System Flow:
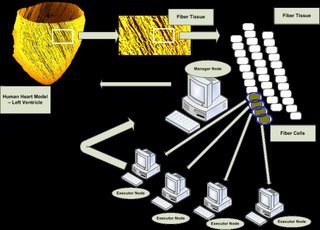
* Construction Data Structure Of Cell Model
* Initialize Timestep For Simulation
* Distribution Cell Computational Tasks to Executor Node(s)
* Computational Tasks of Each Cell Are Executed
* Collect Computational Results from Executors
* Update Computational Results for Cell Model
* Increase Timestep to n+1
More ...

Year: 2004 - 2005
Programming Language & Tools: Programming Language: OpenGL, C, C++, Titanium
Title: Dynamic Cardiac Mechanics Based On Fiber-Fluid Model
Present a non-invasive methodology in constructing a patient-specific virtual human heart
based on fiber-fluid model. I applied digital image processing techniques on patient-specific MR images
for obtaining the geometry information of human heart. The techniques include: image acquisition, image
visualization, image enhancement and segmentation. I incorporated cubic Hermite basis functions in our
epicardium surface interpolation algorithm. I formularized a three-dimensional rule based fiber
reconstruction mechanism to reconstruct the cardiac fiber architecture. A fiber model has been constructed
which consists of 1,038 fibers with 371,239 fiber points. This model describes the ventricular threedimensional
geometry. Immersed Boundary Methods (IBM) is used in constructing fiber-fluid cardiac simulation.
The simulation of early ejection (from 0ms to 0.5ms) for the Left Ventricle (LV) has been implemented in
SGI workstation. Simulation results for cardiac fiber and blood flow are presented in three-dimensional (3D).
Open GL-based animated visualization programs are developed to serve three purposes: (1) to
demonstrate the interpolation and rule-based fiber reconstruction process. (2) To visualize the simulation
results of cardiac fiber and blood flow; (3) To analyse the dynamics of the epicardium fibers as well as the
blood flow in the cavity of the LV.
More ...